First edition 2021 - Directors report
 Dear Colleagues,
Dear Colleagues,
Welcome to the first edition of the newsletter for 2021.
In this issue we highlight some of our recent achievements, new developments and project updates. We are also proud to announce our new collaborative projects for 2021.
The next edition of this newsletter will be distributed in May 2021.
2020 year at a glance
In 2020, GIH was proud to support 20 UQ research projects which spanned across 7 Institutes, 3 Schools and 8 Research facilities, with more than 70 collaborative researchers.
Research supported by our team together with our collaborators resulted in the preparation of 9 manuscripts for publication, 16 grant applications, 3 bioinformatic analysis tools and several global scientific webinars.
The GIH team is excited to continue to drive the success into this new year at UQ with both our existing and new collaborative projects ahead.
Brand new GIH website launched!
 Late last year we were pleased to announce the launch of our new stand alone UQ GIH website by the UQ Pro-Vice Chancellor, Research Infrastructure, Professor Joe Shapter.
Late last year we were pleased to announce the launch of our new stand alone UQ GIH website by the UQ Pro-Vice Chancellor, Research Infrastructure, Professor Joe Shapter.
Please take the time to browse and bookmark our website.
It not only showcases all the novel projects and valuable collaborations GIH is undertaking, but also features information and protocols for the techniques we are developing, links to our publications and bioinformatic software tools, data browsers, online webinars and our news.
Announcing the 2021 GIH collaborative projects
We would like to thank all the UQ researchers who applied for GIH External Projects. We received a number of excellent applications and have had the difficult task of selecting the projects to proceed with this year.
Careful consideration was paid to each projects genomic innovation and feasibility as well as the potential for the developed technology to be accessed by other UQ researchers.
We are pleased to announce the following project collaborations for 2021. The projects selected cover a diverse range of research fields and sample types. They represent both new areas of research for GIH and also build upon our key research strengths.
We look forward to bringing the developments and capabilities from within all these projects to the wider UQ research community soon.
Epigenetics & Epigenomics
Optimising Nanopore Sequencing for Genome-wide DNA Methylation Analysis in Cell-free DNA. Janice Reid, Critical Care Research Group (CCRG). (CCRG is a large research group based at the Prince Charles Hospital and associated with the UQ Faculty of Medicine and the UQ School of Clinical Medicine).
Enabling population scale epigenomics for crop improvement. Peter Crisp, Kai Voss-Fels, Lee Hickey. School of Agriculture and Food Sciences (SAFS) and Queensland Alliance for Agriculture and Food Innovation (QAAFI).
Deciphering the epitranscriptome machinery using direct native RNA sequencing. Cheong Xin Chan, Katherine Dougan, Mikael Bodén, Peter Erskine, Antony van der Ent. Australian Centre for Ecogenomics (ACE), School of Chemistry and Molecular Biology (SCMB), Sustainable Minerals Institute (SMI).
Spatial -omics
Developing new spatial proteomics capability and extending the applications of spatial transcriptomics to clinical archival tissue samples. Quan Nguyen, Mitchell Stark, Kiarash Khosrotehrani, Brett McKinnon. Institute for Molecular Biosciences (IMB) and UQ Diamantina Institute (UQ-DI).
Single-cell (and FACS-based) sequencing technologies
Utilising RNA velocity quantification of single cell transcriptomics to study differential cellular ontogeny in a single stage across different species. Peter Kozulin, Laura Fenlon, Rodrigo Suarez. Queensland Brain Institute (QBI), School of Biomedical Sciences (SBMS).
Expanding the scope of multimodal single cell sequencing. Christian Nefzger. Institute for Molecular Bioscience (IMB).
Nanopore IMPACT: an Integrated Method for Profiling All Cell Types by longread Nanopore sequencing. Paul Marshall, Tim Bredy, Qiongyi Zhao, Martin Smith, Eva Pardo. Queensland Brain Institute (QBI).
Whole genome sequencing
Genome-Phaser: End-to-end protocol for fully phasing whole genome variants using haplotagging. Melanie Wilkinson, Elizabeth Ross, Frank Chan, Craig Hardner and Daniel Ortiz-Barrientos. School of Biological Sciences, QAAFI Centre for Horticultural Science, QAAFI Centre for Animal Science.
Developing a scalable genome browser and interactive repository for large and complex multi-omic datasets from non-model organisms of environmental and economic importance. Sandie Degnan, Bernie Degnan and Dominique Gorse. School of biological sciences (SBS) and Queensland Cyber Infrastructure Foundation (QCIF Bioinformatics).
Culture-independent metagenomic diagnostics for genomic surveillance and infection control of pathogenic bacteria in clinical settings. Brian Forde, Delaney Burnard, David Paterson, Patrick Harris. Centre for Clinical Research (UQCCR).
Assessing DNA Replication Dynamics in Cancer. Mathew Jones and Paul Clarke. UQ Diamantina Institute.
CRISPR & gene editing technologies
High throughput single cell CRISPR technology for experimental validation of population genetics and functional genomics discovery in endometriosis as a model disease. Brett McKinnon and Quan Nguyen. Institute for Molecular Biosciences (IMB).
Optimised bioinformatics and validation pipeline for genome-wide CRISPR screening data. Rebecca San Gil, Adam Walker. Queensland Brain Institute (QBI).
Advancing the sequencing of plant genomes
 The rapid advancement of DNA sequencing technologies has enabled the assembly of highly-accurate, haplotype-resolved plant genomes at the chromosome scale. However, selecting the most suitable and cost-effective method from the range of options available can prove challenging for researchers.
The rapid advancement of DNA sequencing technologies has enabled the assembly of highly-accurate, haplotype-resolved plant genomes at the chromosome scale. However, selecting the most suitable and cost-effective method from the range of options available can prove challenging for researchers.
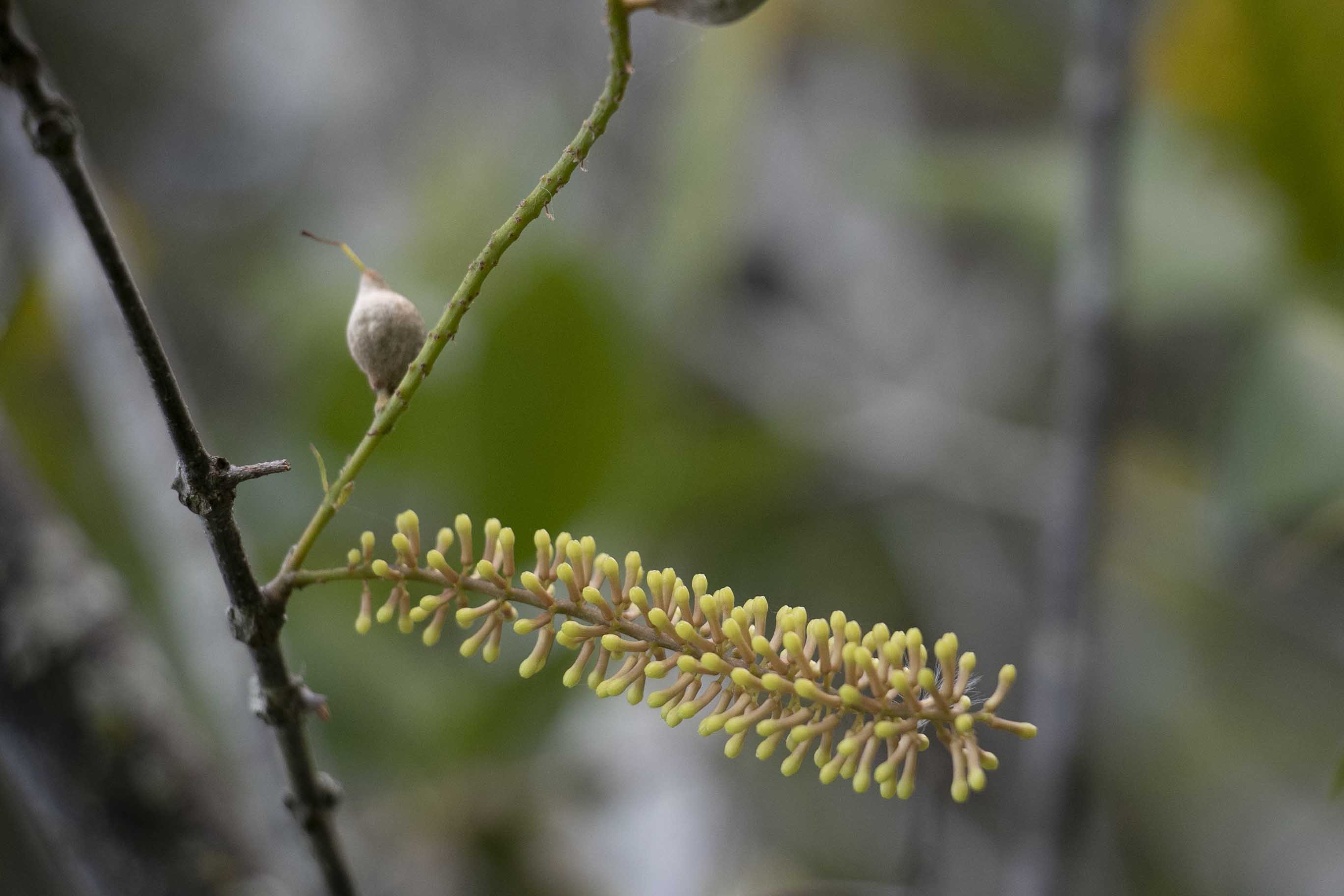
Published recently in the journal GigaScience, our GIH project undertaken in collaboration with a team led by Professor Robert Henry at The Queensland Alliance for Agriculture and Food Innovation (QAAFI), benchmarked 3 long-read sequencing technologies applied to the assembly of the endangered Australian wild Macadamia, Macadamia jansenii. Our optimised plant nuclear HMW DNA extraction for long read sequencing that we used in the study, represented one of our most viewed social media posts with more than 5000 impressions on Twitter.
By utilising cutting-edge sequencing technologies for sequencing some 300 wild and cultivated macadamia varieties, the Hort Innovation National Tree Genomics program is providing genetic insight for growers to improve breeding programs for the nut which has a predicted export value of $350 million by 2025.
Microbial genome web browser launch
 In collaboration with Dr Dom Gorse and the team at QCIF Bioinformatics in a GIH project led by Professor Ian Henderson at the Institute for Molecular Bioscience (IMB), we launched TraDIS-vault, a web-based public repository for visualisation and characterisation of microbial data sets obtained from Transposon Directed Insertion Site Sequencing (TraDIS).
In collaboration with Dr Dom Gorse and the team at QCIF Bioinformatics in a GIH project led by Professor Ian Henderson at the Institute for Molecular Bioscience (IMB), we launched TraDIS-vault, a web-based public repository for visualisation and characterisation of microbial data sets obtained from Transposon Directed Insertion Site Sequencing (TraDIS).
The TraDIS genome sequencing method is commonly used to identify the genes in bacteria that encode essential functions. By making such complex data sets more accessible to researchers, the TraDIS-vault has already begun to host other UQ researchers data sets and there are plans to develop it further to allow submissions from the international community.
Single-cell gene expression in whole tissues
Crowned as the "Method of the Year 2020" by Nature Methods, Spatially resolved transcriptomics, which is the ability to measure gene activity where it occurs, or "spatially", in a whole tissue sense, has revolutionised how researchers can investigate biology and disease.
In a GIH collaboration with the research groups of Dr Quan Nguyen and Professor Andrew Mallett, we have been actively developing Spatial transcriptomics pipelines using the technologies Visium (10X Genomics) and RNAscope (ACD Biosciences).
Also in development as a GIH collaboration, is the technology multiplexed error-robust FISH (MERFISH) with Professor Ernst Wolvetang and his research colleagues. This single-cell genomics technology for > 10,000 genes imaging experiments has been reported with accuracy as high as 80%.
These innovations in the ability to utilise spatial omics technology in exploring how individual cells exist within their tissue microenviroment, though transcriptome analysis is showing great promise in areas such as neuroscience and cancer pathology.
2021 funding success supported by GIH research
 We are thrilled to report some of the recent funding successes received by UQ researchers where GIH developments or capabilities have helped further their research.
We are thrilled to report some of the recent funding successes received by UQ researchers where GIH developments or capabilities have helped further their research.
Dr Mathew Jones and Professor Paul Clarke received an 2021 ARC Discovery Project based on decoding the spatiotemporal control of DNA replication and repair. GIH researchers have been actively collaborating in their research on an innovative method of applying long-read sequencing technology for the analysis of telomeres and DNA replication.
Dr Quan Nguyen with colleagues Dr Laura Genovesi, Prof Brandon Wainwright and A/Prof Marc Ruitenberg were awarded a 2021 NHMRC ideas grant to identify and combat drug resistant cells in paediatric brain cancer using spatial omics. Additionally Quan is part of the team that won the Charcot Prize after being awarded the highest-ranked 2021 MND Research Australia grant. Collaboratively with Quan's excellent team, GIH has established spatial transcriptomics methods and analysis tools using advanced imaging and machine learning techniques.
Dr Adam Walker, Dr Rebecca San Gil and colleagues were selected to lead a 2021 MND Research Australia Innovator grant project utilising new genetic tools to understand the causes of Motor Neurone Disease. As a main part of their research interest, GIH is collaboratively developing a Genome-wide CRISPR screening tool with their research group.
Prof Mark Schembri and his team were awarded a 2021 NHMRC Ideas grant for understanding how antibiotic resistant bacterial superbugs such as E.coli ST131 cause infections. In support of this aim, GIH is undertaking a collaborative project with the research groups of Prof Scott Beatson, Prof Mark Schembri and Prof Mark Walker to develop robust protocols for long read transcriptomics in bacteria, with the test subject E. coli ST131.
GIH collaborator spotlight
 Each newsletter we shine the spotlight on a lead researcher within one of our GIH collaborative projects.
Each newsletter we shine the spotlight on a lead researcher within one of our GIH collaborative projects.
We caught up with Dr Rebecca San Gil, Fight MND Early Career Research Fellow within the Queensland Brain Institute (QBI), who is involved in the GIH project Genome-wide CRISPR screening for modifiers of diverse cellular phenotypes.
What are your key research interests?
I am a passionate researcher intent on dedicating my career to finding new treatments and to cure motor neuron disease and frontotemporal dementia (MND/FTD). With GIH, I am currently using revolutionary gene editing technology to identify new therapeutic targets that combat protein aggregation in neurodegenerative diseases.
Can you tell us a bit about your project with GIH and how this relates to your research?
Our lab with Dr Adam Walker has recently completed an exciting new project using human genome-wide CRISPR screening to identify genes and proteins that regulate protein aggregation in MND/FTD. Aggregation of discrete proteins into protein inclusions is associated with toxicity that causes the death of motor neurons in all cases of MND/FTD. One potential therapeutic strategy is to target and prevent this toxic process of protein clumping, however, the molecular mechanisms involved in driving aggregation remain unclear. This project used revolutionary new gene editing technology to scan all 20,000 genes in the human genome, remarkably this can be achieved in a single experiment. We identified hundreds of potentially disease-modifying genes that may trigger or stop aggregation. This research revealed and fast-tracked the identification of new targets that will be tested for their therapeutic potential in cells and mouse models of MND/FTD.
What is the goal of this project and how could it benefit other researchers?
This technique is highly versatile and will be of interest to those who seek to understand what genes drive fundamental mechanism of action of a cellular pathway or the pathobiology of disease in established cell lines. For example, genome wide CRISPR screens have been applied to understand cancer resistance to chemotherapy drugs, virus virulence factors, protein function and homeostasis, bacterial infection of host cells, immunity and inflammation, and programmed cell death, just to name a few. The technique is also amenable to CRISPR knockout, inhibition or activation. We are willing to collaborate with other groups to adapt the pipeline that we have developed to their own studies.
Find out more about the project >>
 Top tweet
Top tweet
Our latest "top tweet" was regarding our test run for real-time adaptive sampling during long read sequencing on the Oxford Nanopore MinION device.
Adaptive sampling is the ability for a user to electronically select or reject genomic strands of interest in a mixed sample. Traditionally this is done in the sample preparation step and this development represents a fantastic ability for targeted sequencing applications.
Our adaptive sampling is part of the GIH collaborative project "A genomic dissection of metaorganisms: molecular approaches for teasing apart the hologenome". which is led by Dr Cheong Xin Chan, Dr Lauren Messer, Dr Antony Van der Ent, Professor Peter Eskine and their teams from the School of Biological Chemistry and the UQ Centre for Mined Land Rehabilitation, Sustainable Minerals Institute.
With nearly 10,500 impressions and 350 engagements, our post was retweeted many times including by Oxford Nanopore itself.
Discover the adaptive sampling technique >>
Recent GIH developments and protocols
To find out about all our latest developments from within our collaborative projects, click the links below.
Single-cell and short read sequencing
Imaging and spatial transcriptomics
Bioinformatics andcomputational tools
Seminars and Interest groups
Queensland Genomic Technology Innovation Group.
Seminars are every third Tuesday of each month at 2pm.
https://groups.google.com/forum/#!forum/qgti/join
All things up to assembly (atuta.slack.com)
Worldwide Slack channel for discussion regarding sequencing and assembly of genomes.
Nanopore Oceania (nanopore-oceania.slack.com)
Oceania-region based Slack channel to facilitate communication and support between researchers working with Oxford Nanopore-based sequencing, with a strong focus on bioinformatics.
Got news?
We'd love to know if you have found any of our GIH developments helpful for your own research, so drop us a line.
In addition to our quarterly newsletter, GIH news is also posted on our website here and on our Twitter feed here.
Newsletter subscribe. Read all our latest news Visit GIH on Twitter
Copyright © 2012 The University of Queensland
ABN 63 942 912 684 | Privacy policy
Authorised by: Director, Genome Innovation Hub
Maintained by: b.purdue@uq.edu.au